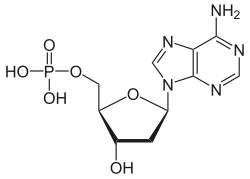
Deoxyadenosine monophosphate
 | |
 | |
| Names | |
|---|---|
| IUPAC name
2′-Deoxyadenylic acid
| |
| Systematic IUPAC name
[(2R,3S,5R)-5-(6-Amino-9H-purin-9-yl)-3-hydroxyoxolan-2-yl]methyl dihydrogen phosphate | |
| Identifiers | |
CAS Number
|
|
3D model (JSmol)
|
|
| ChEBI | |
| ChemSpider | |
| ECHA InfoCard | 100.010.459 |
IUPHAR/BPS
|
|
| MeSH | Deoxyadenosine+monophosphate |
PubChem CID
|
|
| UNII | |
CompTox Dashboard (EPA)
|
|
InChI
| |
SMILES
| |
| Properties | |
Chemical formula
|
C10H14N5O6P |
| Molar mass | 331.222 g/mol |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
Infobox references
| |
Deoxyadenosine monophosphate (dAMP), also known as deoxyadenylic acid or deoxyadenylate in its conjugate acid and conjugate base forms, respectively, is a derivative of the common nucleotide adenosine monophosphate (AMP), in which the -OH (hydroxyl) group on the 2' carbon on the nucleotide's pentose has been reduced to just a hydrogen atom (hence the "deoxy-" part of the name). Deoxyadenosine monophosphate is abbreviated dAMP. It is a monomer used in DNA.
Chemical Properties
The overall structure of dAMP is shaped largely by how its parts interact with each other. In particular, the adenine base and the phosphate group can form hydrogen bond interactions. The distance between the atoms involved in these bonds helps shape the overall conformation of dAMP. The character of these interactions shifts with the molecule’s charge state. When the molecule gains or loses charge, it causes the spacing between the atoms to fluctuate.[1] Increasing the negative charge, tends to draw the atoms closer together, causing the strength of the hydrogen bond to increase as well. [1] Protonation of the adenine increases the strength of the hydrogen bonds. The presence of water molecules can also stabilize the structure by forming additional hydrogen bonds with the phosphate group and the adenine base.[2]
See also
- Nucleic acid
- DNA metabolism
- Cofactor
- Guanosine
- Cyclic AMP (cAMP)
- ATP
References
- ^ a b Shishkin, Oleg V.; Palamarchuk, Gennady V.; Gorb, Leonid; Leszczynski, Jerzy (February 2008). "Opposite charges assisted extra strong C–H⋯O hydrogen bond in protonated 2′-deoxyadenosine monophosphate". Chemical Physics Letters. 452 (1–3): 198–205. doi:10.1016/j.cplett.2007.12.052. Retrieved 8 December 2025.
- ^ Zhang, Mingxing; Zhang, Mingzhen; Liu, Yalan; Chen, Yan; Zhang, Kaiyue; Wang, Chao; Zhao, Xiaoying; Zhou, Chenyang; Gao, Jiawei; Xie, Xinrui; Zheng, Daoyuan; Zhao, Guangjiu (2017-09-01). "DFT/TDDFT theoretical investigation on the excited-state intermolecular hydrogen bonding interactions, photoinduced charge transfer, and vibrational spectroscopic properties of deprotonated deoxyadenosine monophosphate [dAMP–H]− anion in aqueous solution: Upon photoexcitation of hydrogen-bonded model complexes [dAMP–H]−nH2O (n = 0, 1, 2, 3, 4)". Journal of Molecular Liquids. 242: 1118–1122. doi:10.1016/j.molliq.2017.07.120. ISSN 0167-7322.
- Wishart, David S.; Guo, An Chi; Oler, Eponine; Wang, Fel; Anjum, Afia; Peters, Harrison; Dizon, Raynard; Sayeeda, Zinat; Tian, Siyang; Lee, Brian L.; Berjanskii, Mark; Mah, Robert; Yamamoto, Mai; Jovel Castillo, Juan; Torres Calzada, Claudia; Hiebert Giesbrecht, Mickel; Lui, Vicki W.; Varshavi, Dorna; Varshavi, Dorsa; Allen, Dana; Arndt, David; Khetarpal, Nitya; Sivakumaran, Aadhavya; Harford, Karxena; Sanford, Selena; Yee, Kristen; Cao, Xuan; Budinsky, Zachary; Liigand, Jaanus; Zhang, Lun; Zheng, Jiamin; Mandal, Rupasri; Karu, Naama; Dambrova, Maija; Schiöth, Helgi B.; Gautam, Vasuk. "Showing metabocard for Deoxyadenosine monophosphate (HMDB0000905)". Human Metabolome Database, HMDB. 5.0.