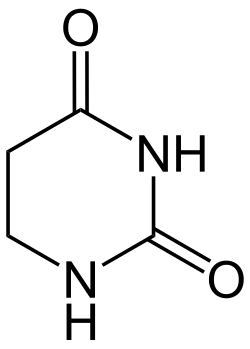
Dihydrouracil
 | |
| Names | |
|---|---|
| Preferred IUPAC name
1,3-Diazinane-2,4-dione | |
| Other names
Dihydropyrimidine-2,4(1H,3H)-dione
| |
| Identifiers | |
CAS Number
|
|
3D model (JSmol)
|
|
| ChemSpider | |
| ECHA InfoCard | 100.007.257 |
| MeSH | Dihydrouracil |
PubChem CID
|
|
| UNII | |
CompTox Dashboard (EPA)
|
|
InChI
| |
SMILES
| |
| Properties | |
Chemical formula
|
C4H6N2O2 |
| Molar mass | 114.10264 |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
Infobox references
| |
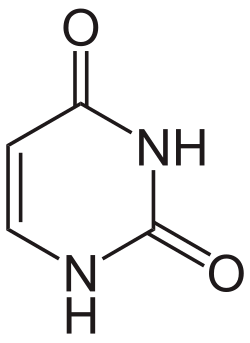
Dihydrouracil is an intermediate in the catabolism of uracil.[1] The enzyme dihydrouracil dehydrogenase (NAD+) converts uracil to dihydrouracil:
The enzyme uses reduced nicotinamide adenine dinucleotide (NADH) as its cofactor.[2][3]
Dihydrouracil is the base present in the nucleoside dihydrouridine.[4]
See also
- Dihydrouracil oxidase
- Dihydropyrimidinase
References
- ^ Nakamura, A; Kikuchi, K; Ohishi, T; Masuike, T (March 2004). "[Assay method for uracil, dihydrouracil, 5-fluorouracil and 5-fluoro-5, 6-dihydrouracil by high-performance liquid chromatography]". Gan to Kagaku Ryoho. Cancer & Chemotherapy. 31 (3): 381–6. PMID 15045945.
- ^ Campbell LL (August 1957). "Reductive degradation of pyrimidines. III. Purification and properties of dihydrouracil dehydrogenase". The Journal of Biological Chemistry. 227 (2): 693–700. doi:10.1016/S0021-9258(18)70749-7. PMID 13462991.
- ^ West, Thomas P. (2001). "Pyrimidine base catabolism in Pseudomonas putida biotype B". Antonie van Leeuwenhoek. 80 (2): 163–167. doi:10.1023/A:1012275512136. PMID 11759049.
- ^ Dalluge, J.; Hashizume, T.; Sopchik, A. E.; McCloskey, J. A.; Davis, D. R. (1996). "Conformational flexibility in RNA: The role of dihydrouridine". Nucleic Acids Research. 24 (6): 1073–1079. doi:10.1093/nar/24.6.1073. PMC 145759. PMID 8604341.