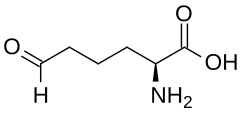
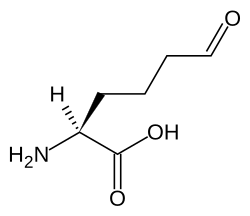
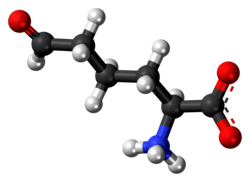
Allysine
 | |
 | |
| Names | |
|---|---|
| Preferred IUPAC name
(2S)-2-Amino-6-oxohexanoic acid | |
| Other names
2-aminoadipate semialdehyde, 2-amino-5-formylvaleric acid, norvaline, 6-oxo-DL-norleucine
| |
| Identifiers | |
CAS Number
|
|
3D model (JSmol)
|
|
| ChEBI | |
| ChemSpider | |
| KEGG | |
| MeSH | allysine |
PubChem CID
|
|
| UNII | |
CompTox Dashboard (EPA)
|
|
InChI
| |
SMILES
| |
| Properties | |
Chemical formula
|
C6H11NO3 |
| Molar mass | 145.158 g·mol−1 |
| Appearance | unstable |
| Density | 1.74g/cm3 |
| Boiling point | 295.2 °C (563.4 °F; 568.3 K) |
| Hazards | |
| Flash point | 132.3 °C (270.1 °F; 405.4 K) |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
Infobox references
| |
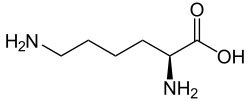
Allysine is a derivative of lysine that features a formyl group in place of the terminal amine. The free amino acid does not exist, but the allysine residue does. It is produced by aerobic oxidation of lysine residues by the enzyme lysyl oxidase. The transformation is an example of a post-translational modification. The semialdehyde form exists in equilibrium with a cyclic derivative.[1]
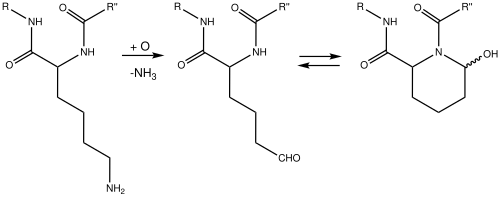
Biochemical reactions
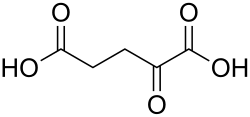
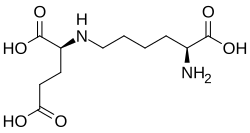
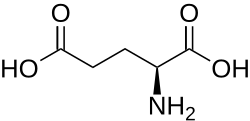
Allysine is linked to L-lysine in reactions catalysed by saccharopine dehydrogenases. These interconvert them via saccharopine:[2][3][4]
Allysine is involved in the production of elastin and collagen.[5] Increased allysine concentration in tissues has been correlated to the presence of fibrosis.[6]
Allysine residues react with sodium 2-naphthol-6-sulfonate to produce a fluorescent bis-naphtol-allysine product.[7] In another assay, allysine-containing proteins are reduced with sodium borohydride to give a peptide containing the 6-hydroxynorleucine (6-hydroxy-2-aminocaproic acid) residue, which (unlike allysine) is stable to proteolysis.[1]
References
- ^ a b Requena, J. R.; Levine, R. L.; Stadtman, E. R. (2003). "Recent Advances in the Analysis of Oxidized Proteins". Amino Acids. 25 (3–4): 221–226. doi:10.1007/s00726-003-0012-1. PMID 14661085. S2CID 28837698.
- ^ Fujioka M, Nakatani Y (1972). "Saccharopine dehydrogenase. Interaction with substrate analogues". Eur. J. Biochem. 25 (2): 301–7. doi:10.1111/j.1432-1033.1972.tb01697.x. PMID 4339117.
- ^ Saunders PP, Broquist HP (1966). "Saccharopine, an intermediate of the aminoadipic acid pathway of lysine biosynthesis. IV. Saccharopine dehydrogenase". J. Biol. Chem. 241 (14): 3435–40. doi:10.1016/S0021-9258(18)96483-5. PMID 4287986.
- ^ Vashishtha AK, West AH, Cook PF (June 2009). "Chemical mechanism of saccharopine reductase from Saccharomyces cerevisiae". Biochemistry. 48 (25): 5899–907. doi:10.1021/bi900599s. PMID 19449898.
- ^ Eyre, David R.; Paz, Mercedes A.; Gallop, Paul M. (1984). "Cross-Linking in Collagen and Elastin". Annual Review of Biochemistry. 53: 717–748. doi:10.1146/annurev.bi.53.070184.003441. PMID 6148038.
- ^ Wahsner J, Désogère P, Abston E, Graham-O'Regan KA, Wang J, Rotile NJ, et al. (April 2019). "68Ga-NODAGA-Indole: An Allysine-Reactive Positron Emission Tomography Probe for Molecular Imaging of Pulmonary Fibrogenesis". Journal of the American Chemical Society. 141 (14): 5593–5596. doi:10.1021/jacs.8b12342. PMC 6494104. PMID 30908032.
- ^ Waghorn PA, Oliveira BL, Jones CM, Tager AM, Caravan P (October 2017). "High sensitivity HPLC method for determination of the allysine concentration in tissue by use of a naphthol derivative". Journal of Chromatography. B, Analytical Technologies in the Biomedical and Life Sciences. 1064: 7–13. doi:10.1016/j.jchromb.2017.08.032. PMC 5662445. PMID 28886479.
Further reading
- Luna C, Estévez M (January 2019). "Formation of allysine in β-lactoglobulin and myofibrillar proteins by glyoxal and methylglyoxal: Impact on water-holding capacity and in vitro digestibility". Food Chemistry. 271: 87–93. doi:10.1016/j.foodchem.2018.07.167. PMID 30236745. S2CID 52309183.
- Luna C, Arjona A, Dueñas C, Estevez M (March 2021). "Allysine and α-Aminoadipic Acid as Markers of the Glyco-Oxidative Damage to Human Serum Albumin under Pathological Glucose Concentrations". Antioxidants. 10 (3): 474. doi:10.3390/antiox10030474. PMC 8002732. PMID 33802856.